Molecule overview
On this page we’ll quickly get an intuitive overview of how Molecule queries and transactions look like.
Pages in the side menu on the right go into more detail.
Db connection
An implicit connection to a database is needed to create and execute molecule queries.
Here’s an example of importing the molecule api and setting up the Datomic Peer in-memory database:
import molecule.datomic.api._
import molecule.datomic.peer.facade.Datomic_Peer._
implicit val conn: Future[Conn] = recreateDbFrom(SomeSchema)
Setup describes the various database setups that can be used with molecules.
Get tuples/objects/json
Molecules fetch data as a Future of tuples, objects or a json String.
Data is retrieved by building molecules of attributes from a namespace. Calling get on a molecule will fetch a Future with tuples of typed data from the database that match the molecule:
val names : Future[List[String]] = Person.name.get
val namesAndAges : Future[List[(String, Int)]] = Person.name.age.get
val namesAgesMembers: Future[List[(String, Int, Boolean)]] = Person.name.age.isMember.get
// etc..
Data can also be returned as an object for each row of data that has properties matching the attributes of the molecule. Note the namespacings ben.Address and ben.Address.City.
for {
// Single row/object
ben <- Person.name_("Ben").age.gender.Address.street.City.name.getObj
_ = ben.age ==> 23
_ = ben.gender ==> "male"
_ = ben.Address.street ==> "Broadway"
_ = ben.Address.City.name ==> "New York"
// Multiple rows/objects
_ <- Person.name.age.Address.street.City.name.getObjList.map { person =>
println(
s"${person.name} is ${person.age} yeas old and lives on " +
s"${person.Address.street}, ${person.Address.City.name}"
)
// "Ben is 23 years old and lives on Broadway, New York"
// "Lisa is ..." etc...
}
} yield ()
Or we can fetch data as a json String with attribute and namespace names used as properties:
for {
_ <- Ns.str.Ref1.int1 insert List(("a", 1), ("b", 2))
_ <- Ns.str.Ref1.int1.getJson.map(_ ==>
"""{
| "data": {
| "Ns": [
| {
| "str": "a",
| "Ref1": {
| "int1": 1
| }
| },
| {
| "str": "b",
| "Ref1": {
| "int1": 2
| }
| }
| ]
| }
|}""".stripMargin
)
} yield ()
The outer structure (“data”) of the json String follows the specification of GraphQL for compatibility.
“CRUD”
In Datomic, data is not deleted but instead “retracted” since all changes are accumulated. That makes it possible to go back and see what data was retracted and is no longer current. That’s why we say “CRUD” instead of CRUD.
// Save populated molecule
Person.name("John").likes("pizza").age(24).save
// Insert multiple tuples of data using a molecule template
Person.name.age.likes insert List(
("John", 24, "pizza"),
("Lisa", 20, "sushi")
)
// Update one or more attributes of a given entity id
Person(johnId).age(25).likes("thai").update
// Retract ("delete") entity
johnId.retract
// Retract attribute value
Person(johnId).likes().update
Attribute types
Card-one attributes
name here is a card-one attribute with a single value
Person.name.get.map(_.head ==> "Bob")
Card-many attributes
interests here is a card-many attribute with a Set of distinct values
Person.name.interests.get.map(_ ==> List(
"Bob", Set("Baseball", "Origami"),
"Liz", Set("Painting", "Traveling", "Tae Kwondo")
))
Map attributes
Keyed card-many attributes, or “Map attributes”, are useful for i18n for instance
for {
_ <- Phrases.greeting("en" -> "hello", "de" -> "hallo").save
_ <- Phrases.greeting("en" -> "hello").get.map(_.head ==> Map("en" -> "hello"))
_ <- Phrases.greeting.k("de").get.ap(_.head ==> Map("de" -> "hallo"))
} yield ()
Mandatory, Optional, Tacit
Attributes can be
- a mandatory value,
- an optional value (
$appended), or - a tacit value (
_appended) that is mandatory but not returned:
// name is mandatory
// age$ is optional
// isMember_ is mandatory but not returned (tacit)
val membersWithOptionalAge: Future[List[(String, Option[Int])]] = Person.name.age$.isMember_.get
Attribute data types
Cardinality one Cardinality many Mapped cardinality many
------------------- ------------------------- --------------------------------
oneString : String manyString : Set[String] mapString : Map[String, String]
oneInt : Int manyInt : Set[Int] mapInt : Map[String, Int]
oneLong : Long manyLong : Set[Long] mapLong : Map[String, Long]
oneDouble : Double manyDouble : Set[Double] mapDouble : Map[String, Double]
oneBigInt : BigInt manyBigInt : Set[BigInt] mapBigInt : Map[String, BigInt]
oneBigDecimal: BigDecimal manyBigDecimal: Set[BigDecimal] mapBigDecimal: Map[String, BigDecimal]
oneBoolean : Boolean manyBoolean : Set[Boolean] mapBoolean : Map[String, Boolean]
oneDate : Date manyDate : Set[Date] mapDate : Map[String, Date]
oneUUID : UUID manyUUID : Set[UUID] mapUUID : Map[String, UUID]
oneURI : URI manyURI : Set[URI] mapURI : Map[String, URI]
oneEnum : String manyEnum : Set[String]
Attribute values
Expressions
// equality
Person.age(42)
// negation
Person.age.not(42) // or
Person.age.!(42)
// comparison
Person.age.>(42)
Person.age.>=(42)
Person.age.<(42)
Person.age.<=(42)
// null
Person.age() // or
Person.age(Nil)
// Word search
Person.comment.contains("nice")
Aggregates
Person.age(count)
Person.age(countDistinct)
Person.age(distinct)
Person.age(max)
Person.age(min)
Person.age(rand)
Person.age(sample)
Person.age(avg)
Person.age(median)
Person.age(stddev)
Person.age(sum)
Person.age(variance)
Logic
OR
Person.age(42 or 43) // same as
Person.age(42, 43) // same as
Person.age(List(42, 43))
AND: card-many attribute category has both a “restaurants” and a “shopping” value:
Community.name.category_("restaurants" and "shopping").get.map(_ ==> List("Ballard Gossip Girl"))
Input molecules
“Input molecules” awaits 1, 2 or 3 input values. Useful for re-use
val personsOfAge = m(Person.name.age_(?))
personsOfAge(23).get.map(_ ==> List("Bob"))
personsOfAge(24).get.map(_ ==> List("Liz", "Don"))
2 inputs + logic (and relationships)
val typeAndRegion = m(Community.name.type_(?).Neighborhood.District.region_(?))
// Social media communities in southern districts
typeAndRegion(("twitter" or "facebook_page") and ("sw" or "s" or "se"))
Relationships
Card-one
Person.name.age.Address.street.get.map(_ ==> List(
("Bob", 23, "5th Avenue")
))
Card-many
// flat
Invoice.no.InvoiceLines.item.get.map(_ ==> List(
(42, "coffee"),
(42, "sugar")
))
// nested
Invoice.no.InvoiceLines.*(InvoiceLine.item).get.map(_ ==> List(
(42, List("coffee", "sugar"))
))
Self-join
Relationship to the same Namespace type (Person -> Person)
Person.name.Spouse.name.get.map(_.head ==> ("Bob", "Liz"))
Directional
Relationships can be defined to go in both directions so that we can traverse a graph uniformly:
Person.name.Knows.name.Knows.name.get.map(_ ==> List(
("Bob", "Liz", "Dan"),
("Dan", "Liz", "Bob")
// etc...
))
Associative
Attributes from different Namespaces that are not explicitly related can be associated by sharing the same entity id. We call molecules with associative relationships “Composite molecules”:
m(Person.name("Bob") + Bar.status("regular")).save
m(Person.name + Bar.status).get.map(_.head ==> ("Bob", "regular"))
Tx Bundle
Transaction bundles can atomically transact multiple operations/statements in one transaction:
transact(
// retract entity
e1.getRetractStmts,
// save new entity
Ns.int(4).getSaveStmts,
// insert multiple new entities
Ns.int.getInsertStmts(List(5, 6)),
// update entity
Ns(e2).int(20).getUpdateStmts
)
Tx meta data
Add meta data to the transaction entity itself about the transaction:
Person.name("John").likes("pizza").Tx(Audit.user("Lisa").uc("survey")).save
We can then query for specific tx meta data
// How was John added?
// John was added by Lisa as part of a survey
Person(johnId).name.Tx(Audit.user.uc).get.map(_ ==> List(("John", "Lisa", "survey")))
// When did Lisa survey John?
Person(johnId).name_.txInstant.Tx(Audit.user_("Lisa").uc_("survey")).get.map(_.head ==> dateX)
// Who were surveyed?
Person.name.Tx(Audit.uc_("survey")).get.map(_ ==> List("John"))
// What did people that Lisa surveyed like?
Person.likes.Tx(Audit.user_("Lisa").uc_("survey")).get.map(_ ==> List("pizza"))
// etc..
Tx function
Ensure transactional atomicity with tx functions that run within a single transaction. If any part of the function throws an exception, the whole transaction is aborted.
Here’s a money transfer function where we want to be sure that both accounts are updated correctly:
// Pass in entity ids of from/to accounts and the amount to be transferred
def transfer(from: Long, to: Long, amount: Int)
(implicit conn: Future[Conn], ec: ExecutionContext): Future[Seq[Statement]] = {
for {
// Validate sufficient funds in from-account
curFromBalance <- Ns(from).int.get.map(_.headOption.getOrElse(0))
_ = if (curFromBalance < amount)
// Throw exception to abort the whole transaction
throw TxFnException(
s"Can't transfer $amount from account $from having a balance of only $curFromBalance."
)
// Calculate new balances
newFromBalance = curFromBalance - amount
newToBalance <- Ns(to).int.get.map(_.headOption.getOrElse(0) + amount)
// Update accounts
newFromStmts <- Ns(from).int(newFromBalance).getUpdateStmts
newToStmts <- Ns(to).int(newToBalance).getUpdateStmts
} yield newFromStmts ++ newToStmts
}
We then call the transaction function inside a transactFn method:
transactFn(transfer(fromAccount, toAccount, okAmount))
Time
Datomic has powerful ways of accessing all the immutable data that accumulates over time in the database:
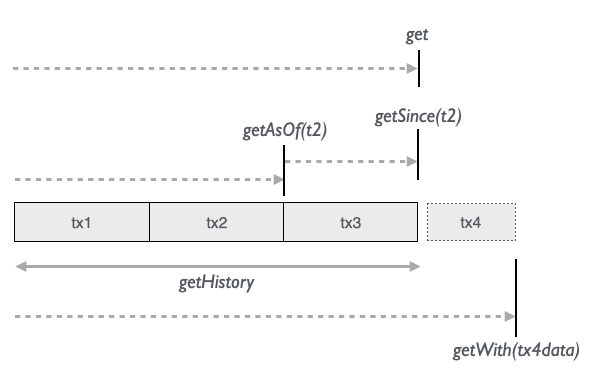
// Current data
Person.name.age.get
// As of some point in time - how did it look at that time?
Person.name.age.getAsOf(nov5date)
// Since some point in time - what has happened after this time?
Person.name.age.getSince(nov5date)
// History of all name operations - what names were added and retracted?
Person.name.getHistory
// Test what-if scenarios given some test statements - how will it look if we do x?
Person.name.getWith(someTestStmts)
Generic APIs
Molecule provides access to Datomic’s various generic interfaces and apis.
Entity API
touch an entity id to get all it’s attribute values
orderId.touch.map(_ ==> Map(
":db/id" -> orderId,
":Order/lineItems" -> List(
Map(
":db/id" -> 102L,
":LineItem/qty" -> 3,
":LineItem/product" -> "Milk",
":LineItem/price" -> 12.0),
Map(
":db/id" -> 103L,
":LineItem/qty" -> 2,
":LineItem/product" -> "Coffee",
":LineItem/price" -> 46.0))
))
Datom
Retrieve generic data about entities and attributes:
// Entity id of Ben with generic Datom attribute `e`
Person.e.name.get.map(_.head ==> (benEntityId, "Ben"))
// When was the Ben's age last changed?
Person.age.txInstant.get.map(_.head ==> (24, <April 4>)) // (Date)
// etc...
Indexes
EAVT
Attributes and values of entity e1
EAVT(e1).a.v.get.map(_ ==> List(
(":Person/name", "Ben"),
(":Person/age", 25),
(":Golf/score", 5.7)
))
AVET
Values, entities and transactions where attribute :Person/age is involved
AVET(":Person/age").e.v.t.get.map(_ ==> List(
(25, e1, t2),
(23, e2, t5)
(14, e3, t7),
))
// AVET index filtered with an attribute name and a range of values
AVET.range(":Person/age", Some(14), Some(24)).v.e.t.get.map(_ ==> List(
(14, e4, t7),
(23, e2, t5)
))
AEVT
Entities, values and transactions where attribute :Person/name is involved
AEVT(":Person/name").e.v.t.get.map(_ ==> List(
(e1, "Ben", t2),
(e2, "Liz", t5)
))
VAET
Get entities pointing to entity a1
VAET(a1).v.a.e.get.map(_ ==> List(
(a1, ":Release/artists", r1),
(a1, ":Release/artists", r2),
(a1, ":Release/artists", r3)
))
Log
Data from transaction t1 until t4 (exclusive)
Log(Some(t1), Some(t4)).t.e.a.v.op.get.map(_ ==> List(
(t1, e1, ":Person/name", "Ben", true),
(t1, e1, ":Person/age", 25, true),
(t2, e2, ":Person/name", "Liz", true),
(t2, e2, ":Person/age", 23, true),
(t3, e1, ":Person/age", 25, false),
(t3, e1, ":Person/age", 26, true)
))
Schema
Get information about the Datomic schema which is also a way to access your Data Model programatically.
Schema.part.ns.attr.fulltext$.doc.get.map(_ ==> List(
("ind", "Person", "name", Some(true), "Person name"), // fulltext search enabled
("ind", "Person", "age", None, "Person age"),
("cat", "Sport", "name", None, "Sport category name")
))